MarkerLab®
Omics data organized and presented in an intuitive, engaging and enlightening format.
MarkerLab is a web-based data organization and visualization platform that helps labs publish rich interactive reports to researchers across a wide range of applications.
We developed MarkerLab to enhance core lab capabilities; with simple tools for data organization, a flexible knowledge base aligned to lab-specific offerings, and rich interactive reports that make lab data more accessible, interpretable and valuable to research customers.
For Labs
The MarkerLab platform enables custom strategies for organizing and publishing multi-omics data; with accompanying support for marketing and selling data services, and extensive integration with third-party kits, methods and platforms to enhance your offering and provide optimum value for your research customers.
For Kit and Reagent Companies
MarkerLab provides an extensible framework for promoting and delivering complete kit-to-presentation solutions to labs, and a targeted distribution network for reaching the larger research community.
For Researchers
MarkerLab presents a delightful user experience with beautiful, intuitive tools for organizing, visualizing and understanding multi-omic data, and collaborating with colleagues.
Sharing and collaborating on omics data has never been easier!
Focused on usability and putting data in the right biological or chemical context, MarkerLab makes it easy to load, visualize and learn from multi-omics data.
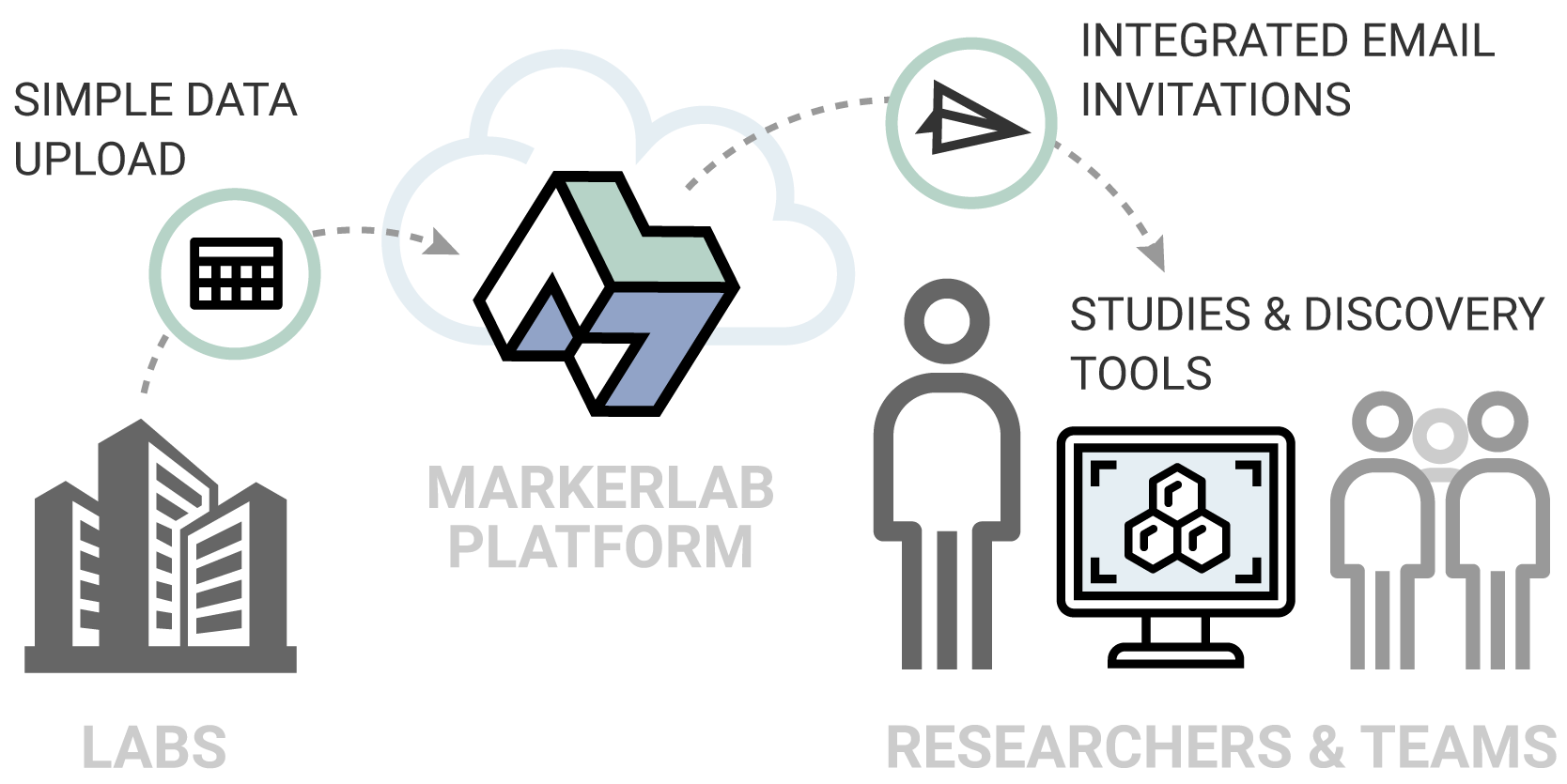
MarkerLab enables labs to publish data directly to research customers, and enables customers to establish collaborative teams that work together in context with their study data.
MarkerLab Apps provide an integrated collection of data organizing tools including marker annotations, reference links, marker ontologies, equations, and pathway maps. These tools are easily modified to produce a custom reporting environment where data and interpretive content are effectively organized to support lab-specific offerings and applications.
MarkerLab By Design

EASY DATA UPLOAD
Drag and drop to upload study data in the prescribed CSV format, and include additional study reference and supporting documentation in PDF, DOCX, PPTX, and XLS. All study files remain conveniently accessible and available for download at any time.
DYNAMIC MARKER MATCHING
MarkerLab's proprietary matching algorithm associates individual marker names with corresponding public database IDs. The accompanying editor enables additional matching and modifications based on lab-specific preferences.

CUSTOMIZABLE APPS
MarkerLab Apps provide essential tools for organizing data into interpretable structures. Biomarker sets, annotations, equations, and pathway maps can be created, edited and applied across the platform—allowing each user to define and distribute unique strategies for organizing data.

SIMPLE SHARING
MarkerLab provides a simple web form with integrated email for sharing studies within the platform, or inviting new users to sign up and collaborate. MarkerLab makes it easy for study owners to review the list of study participants and assign, modify, or remove user permissions for any study at any time.
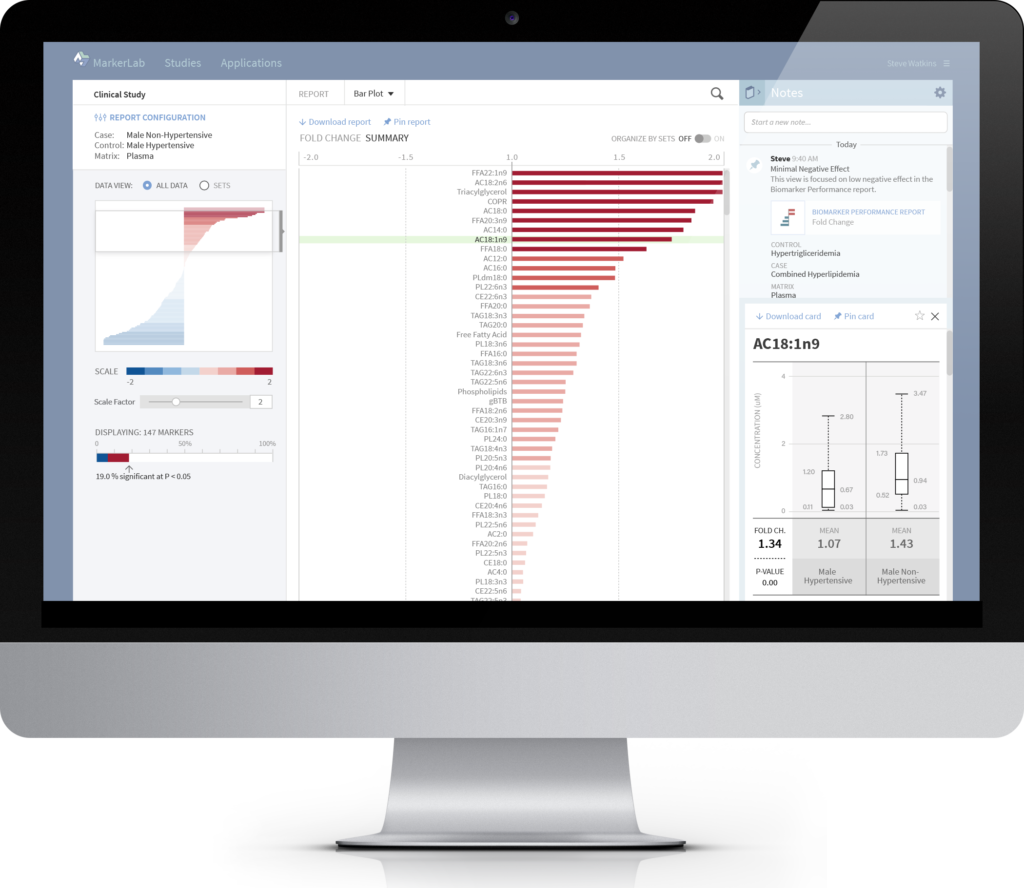
INTERACTIVE VISUALIZATION
MarkerLab's interactive reporting environment provides intuitive controls that enable researchers to create optimal data visualizations, save and recall specific report settings, search and highlight markers of interest, record findings, and form hypotheses for future experiments or deeper statistical analysis.

DOCUMENTATION & COLLABORATION
Integrated notebook functionality enables researchers to maintain a dynamic collection of study notes, recorded in context with data visualizations. Notes are recorded in chronological order and can be shared between study collaborators, including bookmarked links to specific report views for convenient reference.
MarkerLab Apps: Your Data In Context
MarkerLab applications allow you to define your own data organization strategies, and organize your study data into custom, interpretable structures.
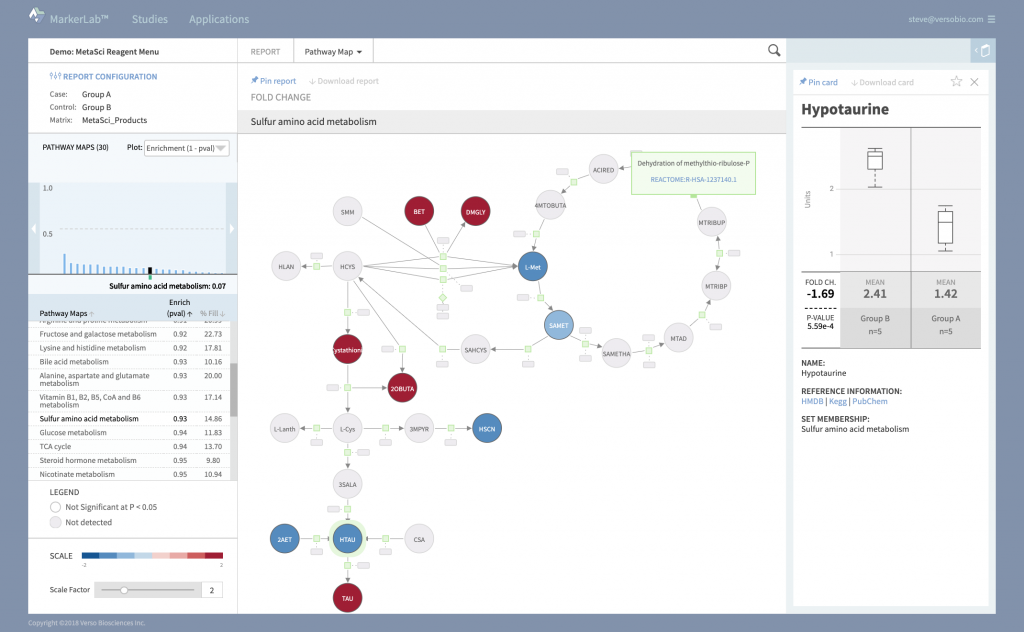
Pathways
MarkerLab includes an extensive collection of metabolic and gene/protein network pathways, curated from publicly available resources including reactome.org. Pathway tools provide authoring, editing, and import functionality so you can customize or add new maps that put your data in the best possible context.

Marker Sets & Ontologies
Sets and ontologies enable critical signal detection and enrichment methods—key to the underlying data organization approach in MarkerLab. These tools include reference to a comprehensive collection of default marker sets derived from reactome.org, with the capability to create, edit, and share your data organization strategies across the platform.
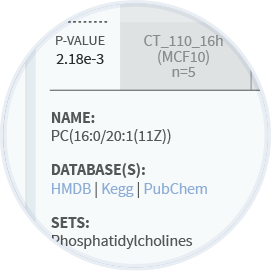
Annotations
MarkerLab provides annotations for individual biomarkers in context with every report and across all studies. Default annotations are mapped to database IDs from a wide variety of preferred resources such as Uniprot, Ensemble, Pubchem, HMDB and KEGG. In addition, the annotations tool makes it easy to edit individual references or upload new content in bulk from external files.
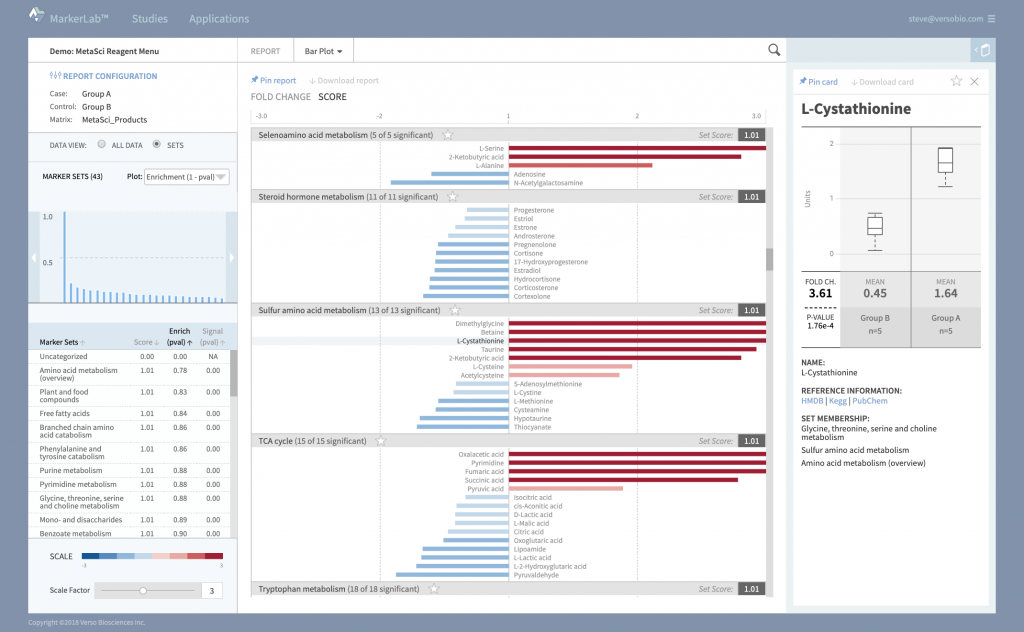
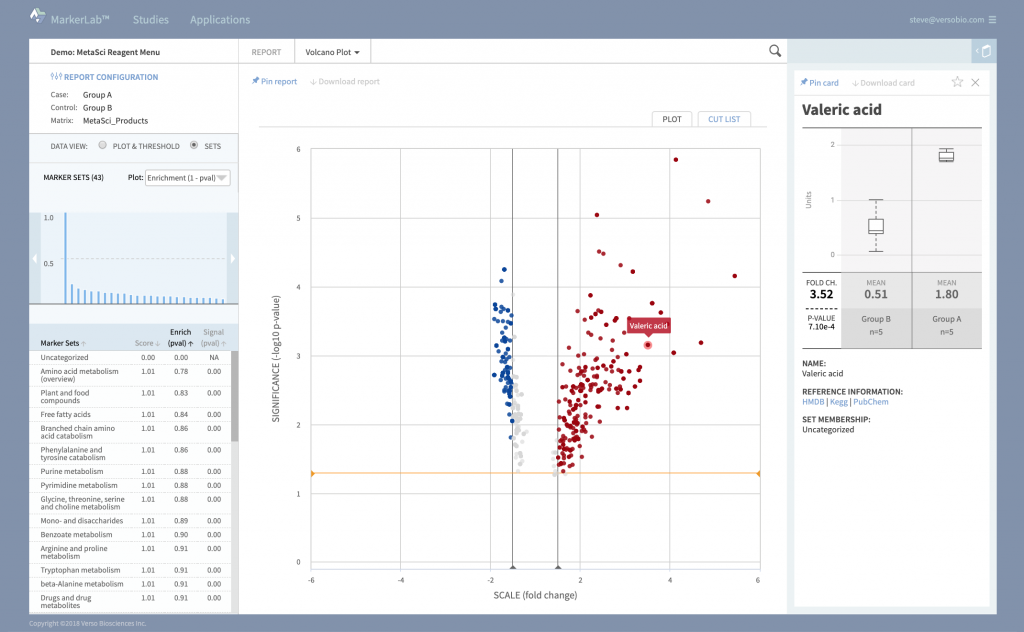
Beautiful Interactive Reports
Dynamic reports provide multiple levels of data interaction and evaluation.
Review experimental effects across the entire data set, at the pathway and marker set level, or in detailed data cards that describe specific effects on individual markers.